CLARITY whole-brain segmentation
There are multiple segmentation functions for different data (stains/channels):
viruscFossparsenuclear
The segmentation workflow relies on an output from the registration workflow, but the segmentation wrapper function can be run without running the registration workflow.
This workflow performs the following tasks:
Segments neurons in cleared mouse brain of sparse or nuclear stains in 3D
Voxelizes segmentation results into density maps with Allen Atlas resolution
Computes features of segmented image and summarizes them per label
It executes:
seg/miracl_seg_clarity_neurons_wrapper.sh
seg/miracl_seg_voxelize_parallel.py
seg/miracl_seg_feat_extract.py
Main outputs
File |
Description |
|---|---|
|
Segmentation image with all labels (cells) |
|
Binarized segmentation image |
|
Segmentation results voxelized to ARA resolution |
|
Binarized version |
|
Segmentation features summarized per ARA labels |
Hint
Results can be opened in Fiji for visualization
GUI
Select from the main GUI menu (invoked from the cli: $ miraclGUI) or run:
$ miracl flow seg
The following window will appear:

Click on Select registered labels (..clar_vox.tif) in the reg_final dir
to choose the registered labels
annotation_hemi_{side}_**um_clar_vox.tif to summarize segmentation
features where:
{side}->combinedorsplit**is the resolution ->10,25or50
The following window will appear:

Next, click on select input tiff dir to select folder with Thy1-YFP
or other channel:
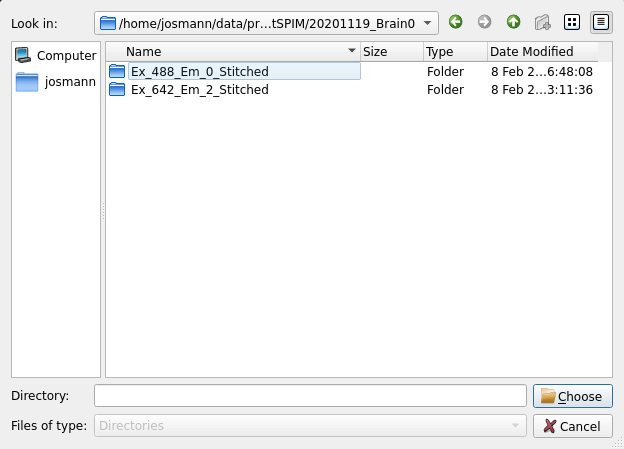
Lastly set the segmentation parameters:
Parameter |
Description |
Default |
|---|---|---|
seg type |
Channel type:
|
|
channel prefix |
Channel prefix and number if multiple channels. Example: |
|
labels voxel size |
Registered labels voxel size in um:
|
|
Click Enter and Run to start the segmentation process.
Command-line
Usage:
$ miracl flow seg -f [ Tiff_folder ]
Example:
$ miracl flow seg -f my_tifs -t nuclear -s "-p C001" -e "-l reg_final/annotation_hemi_combined_25um_clar_vox.tif"
Arguments:
arguments (required):
f. Input Clarity tif folder/dir [folder name without spaces]
t. Channel type: sparse (like Thy1 YFP) or nuclear (like PI)
optional arguments (don't forget the quotes):
Segmentation (invoked by -s " "):
p. Channel prefix & number if multiple channels (like Filter0001)
Feature extraction (invoked by -e " "):
l. Allen labels (registered to clarity) used to summarize features
reg_final/annotation_hemi_{hemi}_{vox}um_clar_vox.tif